FragPipe:
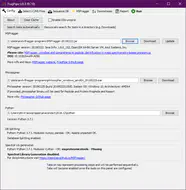
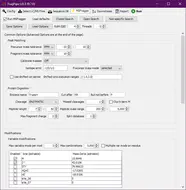
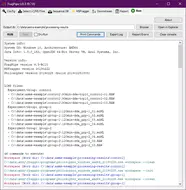
FragPipe is a Java Graphical User Interface (GUI) for a suite of computational tools enabling comprehensive analysis of mass spectrometry-based proteomics data. It is powered by MSFragger - an ultrafast proteomic search engine suitable for both conventional and “open” (wide precursor mass tolerance) peptide identification. FragPipe also includes the Philosopher toolkit for downstream post-processing of MSFragger search results (PeptideProphet, iProphet, ProteinProphet), FDR filtering, label-free quantification, and multi-experiment summary report generation. Crystal-C and PTM-Shepherd are included to aid interpretation of open search results. Also included in FragPipe binary are SpectraST-based spectral library building module, and DIA-Umpire SE module for direct analysis of data independent acquisition (DIA) data.